Aspergillus oryzae and flavus: Difference between revisions
Jump to navigation
Jump to search
No edit summary |
No edit summary |
||
| (3 intermediate revisions by the same user not shown) | |||
| Line 9: | Line 9: | ||
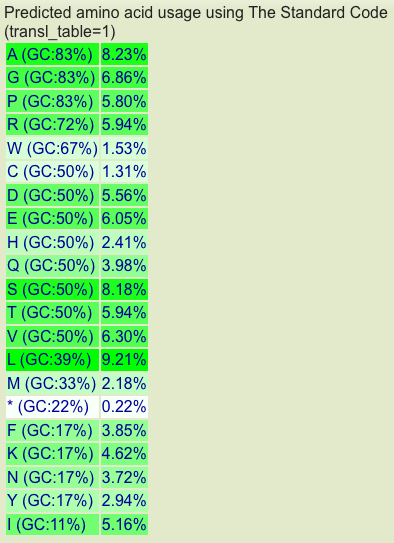
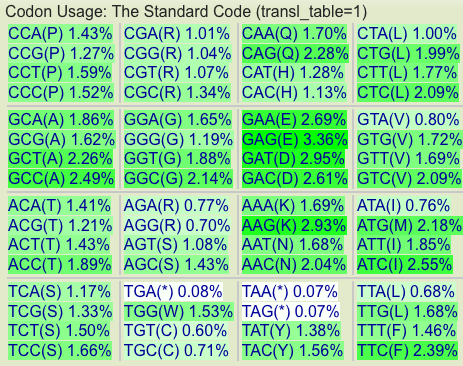
[[Image:Aspergillua-oryzae-amino-acid-usage-table.png|thumb|600px|right|Aspergillus oryzae amino acid usage table for all protein coding sequences.]] | [[Image:Aspergillua-oryzae-amino-acid-usage-table.png|thumb|600px|right|Aspergillus oryzae amino acid usage table for all protein coding sequences.]] | ||
Overview of genomes: | ==Overview of genomes:== | ||
*Aspergillus oryzae strain RIB 40: http://genomevolution.org/CoGe/OrganismView.pl?dsgid=8171 | *Aspergillus oryzae strain RIB 40: http://genomevolution.org/CoGe/OrganismView.pl?dsgid=8171 | ||
*Aspergillus flavus strain NRRL3357: http://genomevolution.org/CoGe/OrganismView.pl?dsgid=7146 | *Aspergillus flavus strain NRRL3357: http://genomevolution.org/CoGe/OrganismView.pl?dsgid=7146 | ||
==Secondary metabolic gene clusters== | |||
[[Aspergillus flavus secondary metabolic gene clusters]] | |||
Latest revision as of 20:08, 27 April 2010



Overview of genomes:
- Aspergillus oryzae strain RIB 40: http://genomevolution.org/CoGe/OrganismView.pl?dsgid=8171
- Aspergillus flavus strain NRRL3357: http://genomevolution.org/CoGe/OrganismView.pl?dsgid=7146