Ancestral angiosperm genomes: Difference between revisions
Jump to navigation
Jump to search
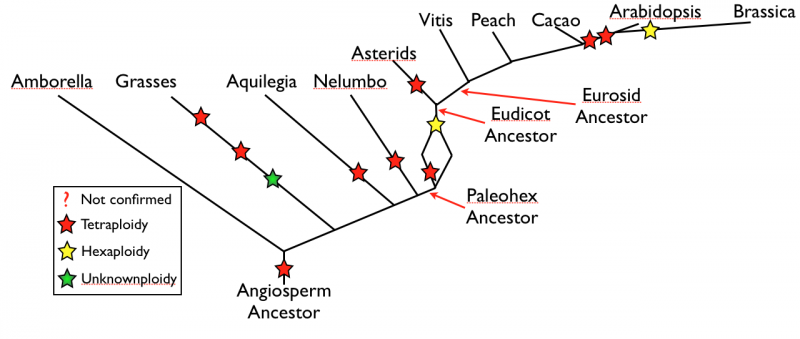
Created page with 'Phylogeny of the Angiosperms with polyploidy events marked. Branch lengths are meaningless.' |
|||
| (5 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
[[File:Screen shot 2012-06-18 at 10.51.11 AM.png|Phylogeny of the Angiosperms with polyploidy events marked. Branch lengths are meaningless.]] | [[File:Screen shot 2012-06-18 at 10.51.11 AM.png|thumb|center|800px|Phylogeny of the Angiosperms with polyploidy events marked. Branch lengths are meaningless.]] | ||
==Ancestral Eurosid== | |||
# Data generated by Chungfang Zheng and David Sankoff | |||
# Procedure for generating ancestral genome: | |||
## Take ancestral ortholog sets from grape/chocolate/peach | |||
## Use the ortholog gene from first grape, then chocolate, then peach (if data is missing) | |||
## Extract CDS sequence from CoGe | |||
## Reverse complement if necessary | |||
## Print sequence | |||
## Pad with 1000 "N"s between coding sequence | |||
Link to ancestral genome: http://genomevolution.org/CoGe/OrganismView.pl?oid=38348 | |||
===Syntenic dotplots=== | |||
[[Eurosid Ancestor to Eurosid Ancestor]] | |||
[[Eurosid Ancestor to Grape]] | |||
[[Eurosid Ancestor to Cacao]] | |||
[[Eurosid Ancestor to Peach]] | |||
[[Eurosid Ancestor to Amborella]] | |||
==Pre-hexaploid eudicot ancestor== | |||
# Data generated by Chungfang Zheng and David Sankoff | |||
# Procedure for generating ancestral genome: | |||
## Take ancestral ortholog sets from grape/chocolate/peach | |||
## Use the ortholog gene from first grape, then chocolate, then peach (if data is missing) | |||
## Extract CDS sequence from CoGe | |||
## Reverse complement if necessary | |||
## Print sequence | |||
## Pad with 1000 "N"s between coding sequence | |||
[[Paleohex Ancestor to Paleohex Ancestor]] | |||
[[Paleohex Ancestor to Grape]] | |||
[[Paleohex Ancestor to Cacao]] | |||
[[Paleohex Ancestor to Peach]] | |||
[[Paleohex Ancestor to Nelumbo]] | |||
[[Paleohex Ancestor to Amborella]] | |||
Latest revision as of 17:43, 19 June 2012
Ancestral Eurosid
- Data generated by Chungfang Zheng and David Sankoff
- Procedure for generating ancestral genome:
- Take ancestral ortholog sets from grape/chocolate/peach
- Use the ortholog gene from first grape, then chocolate, then peach (if data is missing)
- Extract CDS sequence from CoGe
- Reverse complement if necessary
- Print sequence
- Pad with 1000 "N"s between coding sequence
Link to ancestral genome: http://genomevolution.org/CoGe/OrganismView.pl?oid=38348
Syntenic dotplots
Eurosid Ancestor to Eurosid Ancestor
Pre-hexaploid eudicot ancestor
- Data generated by Chungfang Zheng and David Sankoff
- Procedure for generating ancestral genome:
- Take ancestral ortholog sets from grape/chocolate/peach
- Use the ortholog gene from first grape, then chocolate, then peach (if data is missing)
- Extract CDS sequence from CoGe
- Reverse complement if necessary
- Print sequence
- Pad with 1000 "N"s between coding sequence