Fractionation Mutagenesis
What is fractionation mutagenesis?
Natural promoter bashing is a technique for developing testable hypothesis about the function of promoter regions or conserved promoter elements by taking advantage of fractionation mutagenesis using a combination of comparative genomics and comparative expression studies.
Whole genome duplications create two copies of every gene in a genome, each copy with identical promoters containing identical regulatory elements. We already know that duplicate copies of many genes are lost following whole genome duplications, usually by short to medium sized deletions. Genomes of species like maize, where a whole genome duplication occurred 5-12 million years ago, still contain gene fragments that show evidence of "bites" taken out of them. (Should I show an example of this?)
Deletions are not confined to the coding regions of genes but can also remove regions of upstream regulatory sequence.
Natural promoter bashing starts by identifying duplicate genes which show dissimilar patterns of expression with regards to some criteria a researcher is interested in. Perhaps one copy of the gene is expressed only in a certain cell type and the other is not. Or one gene is upregulated in response to a stimulus like drought stress and the other is not. Or one gene shows a change of expression in a mutant background and the other does not.
It is important that the difference observed is a difference in pattern of expression rather than absolute level of expression. Whole genome duplicates can show identical patterns of expression while being expressed at very different absolute levels. It is thought that these differences are mediated by chromatin environment rather than specific deletions/insertions in promoters of one gene or the other.
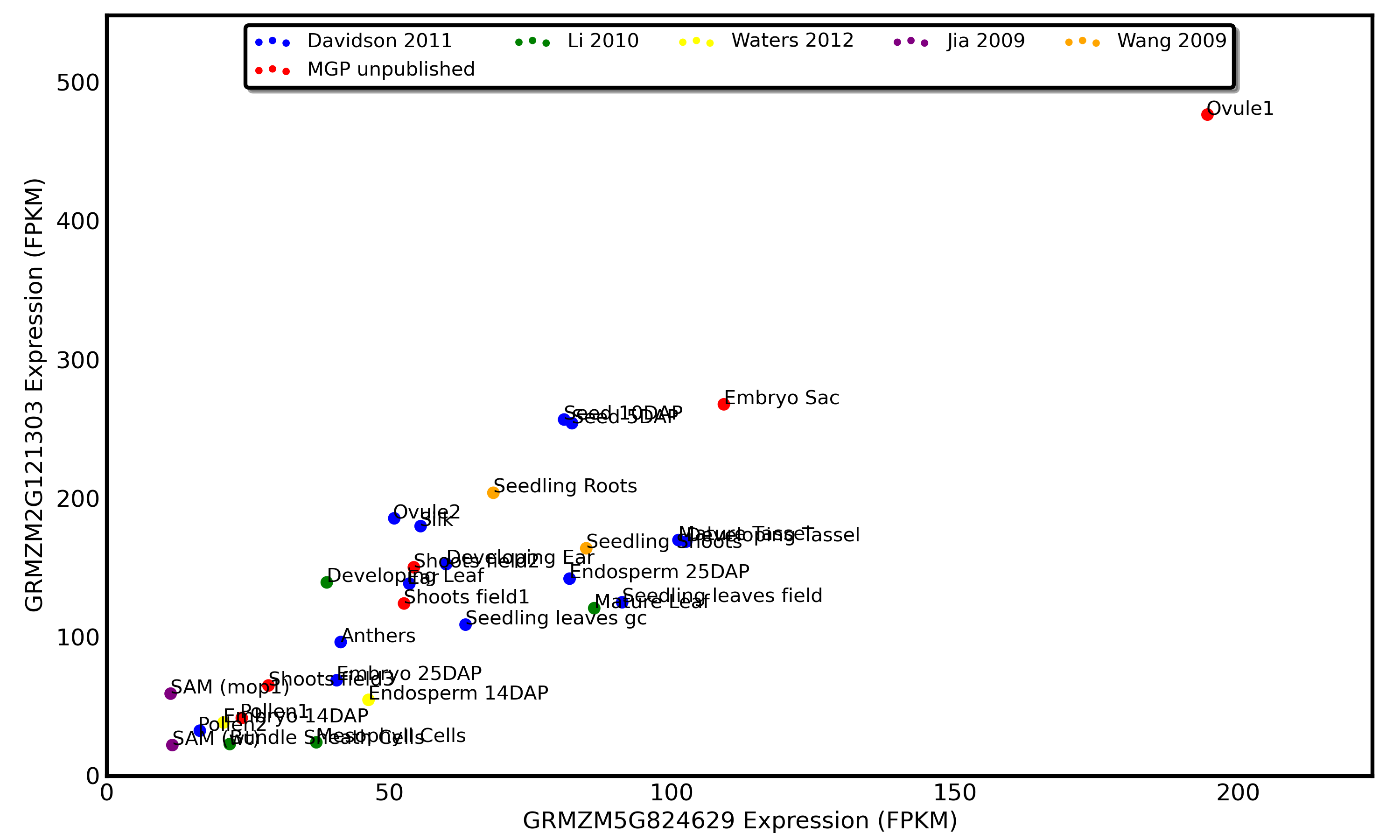
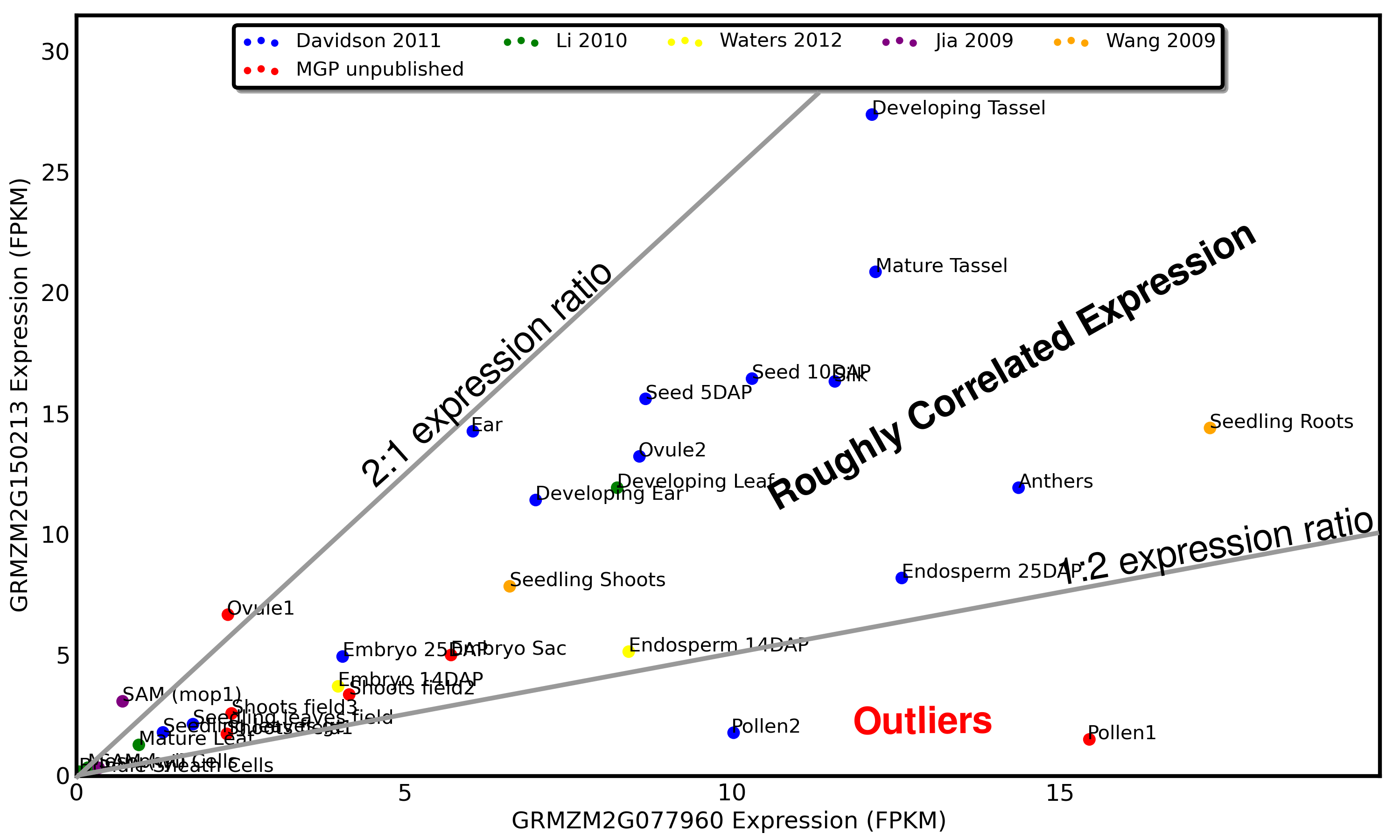
For this example we will use a duplicate pair of genes in maize which show a very different pattern of expression in endosperm relative to other tissues.
Note that the exceptional ratio of expression of this gene pair in developing embryos is supported by two different datapoints from two different research groups (Waters 2012 and Davidson 2012).
Based on this pattern of expression we can hypothesize that GRMZM2G085049 (x-axis) has lost a embryo specific enhancer or GRMZM2G012814 has lost an embryo specific repressor. Both of the models proposed above make assumption that a a regulatory sequence has been LOST from one gene rather than one gene gaining a new piece of regulatory DNA. While loss of function mutations should be much more common than gain of function ones, check out the "Gotchas to Look Out For" section below.
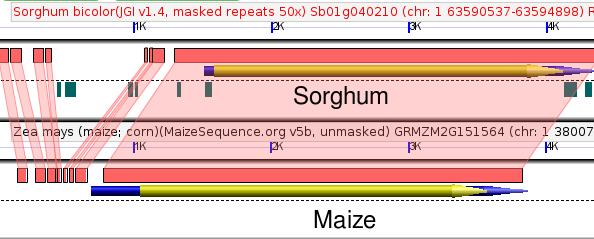
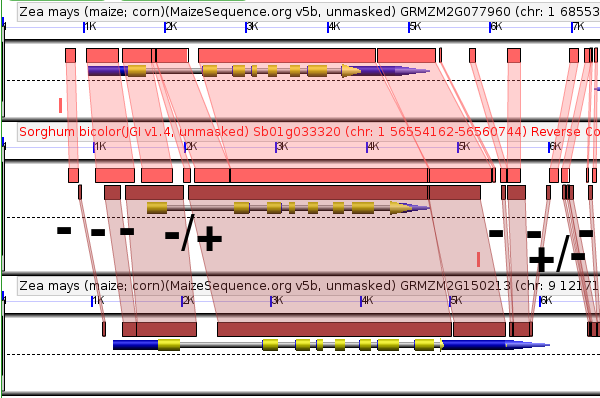
To track changes in the promoter sequence surrounding these two genes we can compare both genes to their shared sorghum ortholog. Sorghum diverged from maize around the same time as the maize whole genome duplication, so even functionless sequence should still show some detectable similarity between sorghum and maize (assuming it hasn't been deleted).
As you can see there are several sequences conserved between GRMZM2G012814 and sorghum but missing from GRMZM2G085049 and vice versa. But how to we prioritize those sequences?