LoadExperiment
This page is currently being updated to reflect the new LoadExperiment version deployed on March 24th.
LoadExperiment enables you to load a file of experimental quantitative, polymorphism, or alignment data for a genome in CoGe. Several different file formats are supported. The data can then be viewed alongside annotation in GenomeView.
Inputs
Metadata
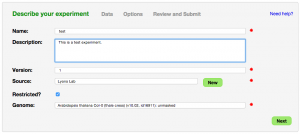
- Name: Name of experiment
- Description: Description of experiment
- Version: Version of experiment
- Source: Where is the data from? This could be you, your lab, your university, a sequencing center, your collaborator.
- Restricted: Is this experiment public or restricted to you and your collaborators
- Genome: Select the appropriate genome from CoGe to associate file with
- Select Data File: Opens a window for specifying the input data file
- Note: Additional metadata about the experiment can be added as well.
- Example from an experiment loaded into EPIC-CoGe: http://genomevolution.org/CoGe/ExperimentView.pl?eid=193
- Information on providing a metadata file for bulk import: Experiment Metadata
Data Files
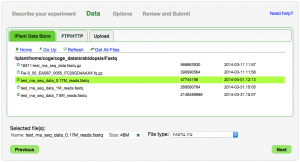
You can select and retrieve data file located at:
- The iPlant Data Store
- An FTP server
- Your computer (Upload)
Note: Typically only a single data file can be selected. The exception is FASTQ formatted files for which multiple files can be selected.
Data Formats and Track Types
LoadExperiment supports several data file formats depending on the data type:
- Quantitative data
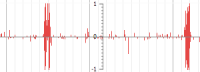
Quantitative track - Comma-separated (CSV) file format
- Tab-separated (TSV) file format
- BED file format
- WIG file format
- Marker data

Marker track - GFF/GTF file format
- Polymorphism (SNP) data

SNP track - Variant Call Format (VCF) file format
- Alignment data
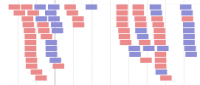
Alignment track - BAM file format
Each of these file formats are described below in their own section. The file type can be auto-detected by LoadExperiment if the file name ends with the expected extension (.csv, .tsv, .bed, .gff, .gtf, .vcf, .bam). Files can be compressed (.zip, .gz) and still have their type auto-detected (e.g., mydata.bed.gz). For non-standard file name extensions, you can select the file type from a list.
CSV File Format
This is a comma-delimited file that contains the following columns
- Chromosome (string)
- Start position (integer)
- Stop position (integer)
- Chromosome Strand (1 or -1)
- Measurement Value must be between [0-1] (real number; inclusive)
- Second Value (OPTIONAL): can store a second value such as an expect value (real number)
#CHR,START,STOP,STRAND,VALUE1(0-1),VALUE2(ANY-ANY) Chr1,11486,12316,1,0.181430277220112,7.3980806218146 Chr1,27309,28272,1,0.944373742485446,5.08225285439412 Chr1,32484,32978,1,0.328500324191726,1.97719838086201 Chr1,41942,42508,-1,0.825027233105203,6.56057592312617 Chr1,56394,57527,-1,0.183234367788511,0.795527328556531 Chr1,67705,68809,-1,0.956523086778851,5.20992343466606 Chr1,71144,72409,1,0.42955128220331,1.80604269639474 Chr1,81671,82833,1,0.626003507696723,2.77834108023821 Chr1,86467,87623,-1,0.0878653961575928,7.42843749315945
TSV File Format
Same as CSV format but with tab delimiters instead of commas.
BED File Format
Two types are supported:
- Standard BED format as defined here: http://genome.ucsc.edu/FAQ/FAQformat.html#format1
- Only the first six columns are used, with the "name" field ignored.
- BedGraph format as defined here: http://genome.ucsc.edu/goldenPath/help/bedgraph.html
Note that the 0-based coordinates of the BED format will be translated into 1-based within CoGe. Measurement values must be between [0,1].
WIG File Format
Standard WIG format as defined here: http://genome.ucsc.edu/goldenpath/help/wiggle.html
Only the "variableStep" type is supported. The "fixedStep" type is not supported and will cause the load to fail with an error.
Start and end coordinates are expected to be 1-based. Measurement values must be between [0,1].
GFF File Format
Standard GFF3 format as defined here: http://gmod.org/wiki/GFF3
Only the seqid, start, end, score, strand, and attribute columns are used (column numbers 1, 4, 5, 6, 7, 9 respectively).
VCF File Format
Standard VCF 4.1 format as defined here: http://www.1000genomes.org/wiki/Analysis/Variant%20Call%20Format/vcf-variant-call-format-version-41
BAM File Format
Standard BAM format.
FASTQ Data
EPIC-CoGe now supports fastq data generated by RNASeq. When loaded, EPIC-CoGe will run and the Expression Analysis Pipeline developed by James Schnable for his qTeller project.
Troubleshooting
Make sure that these basic requirements are followed:
- The file format must be one of those listed above.
- The chromosome names used in the data files exactly match those for the genome. Mismatches will cause the load to fail.
- For text files, such as .CSV and .VCF, the newline (EOL) characters must be Unix-compliant (LF character, not CRLF) , see this article. This is sometimes a problem when loading text files created in Excel on Windows OS.
Bulk Loading
Please contact the CoGe Team if you have large numbers of experiments you wish to load and we can help you with the bulk loading.