Genomic Rearrangement Analysis
From CoGepedia
Contents
Tutorial:
Using CoGe to help with Genomic Rearragement Analyses is quite simple:
- Pre-configured SynMap example: http://genomevolution.org/r/2nd9
- Go to SynMap
- Select two genomes
- Select the "Analysis Options" tab
- Optional: Set the "Merge Syntenic Blocks" algorithm to [[Quota align] and set some number of allowable overlapping genes (10-20 works well for microbes, select 80 for large animal and plant genomes)
- Set the "Syntenic Depth" algorithm to Quota align and have the ratio of syntenic depth set to 1:1
- Run SynMap
- Use the link "Rearrangement Analysis" located under the "Links and Downloads". This will take you to GRIMM for the genomic rearrangement analysis
- Run GRIMM
Configuring Quota Align with a 1:1 ratio of syntenic depth to auto-enable the link to GRIMM for genomic rearrangement analysis
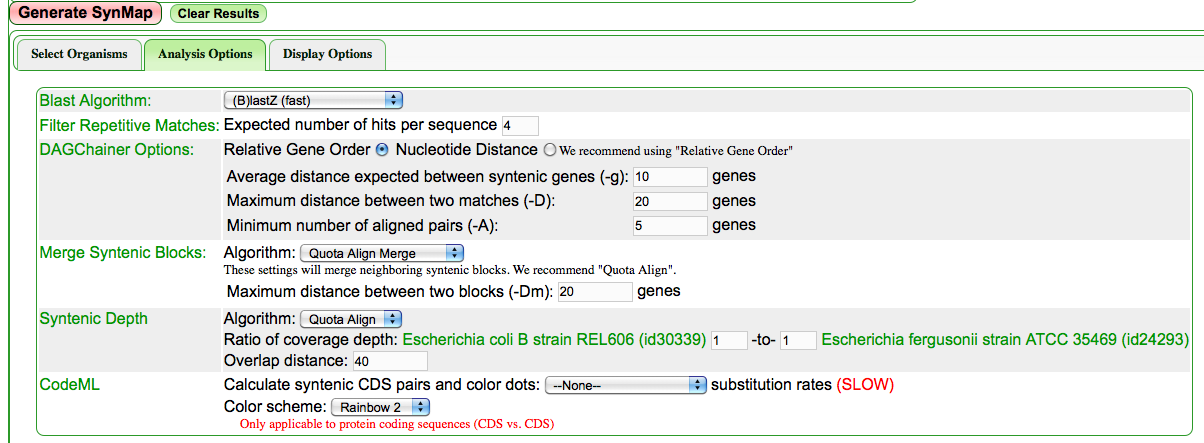
Configuring a SynMap analysis to use Quota align with a syntenic depth of 1:1.
SynMap results with link to GRIMM
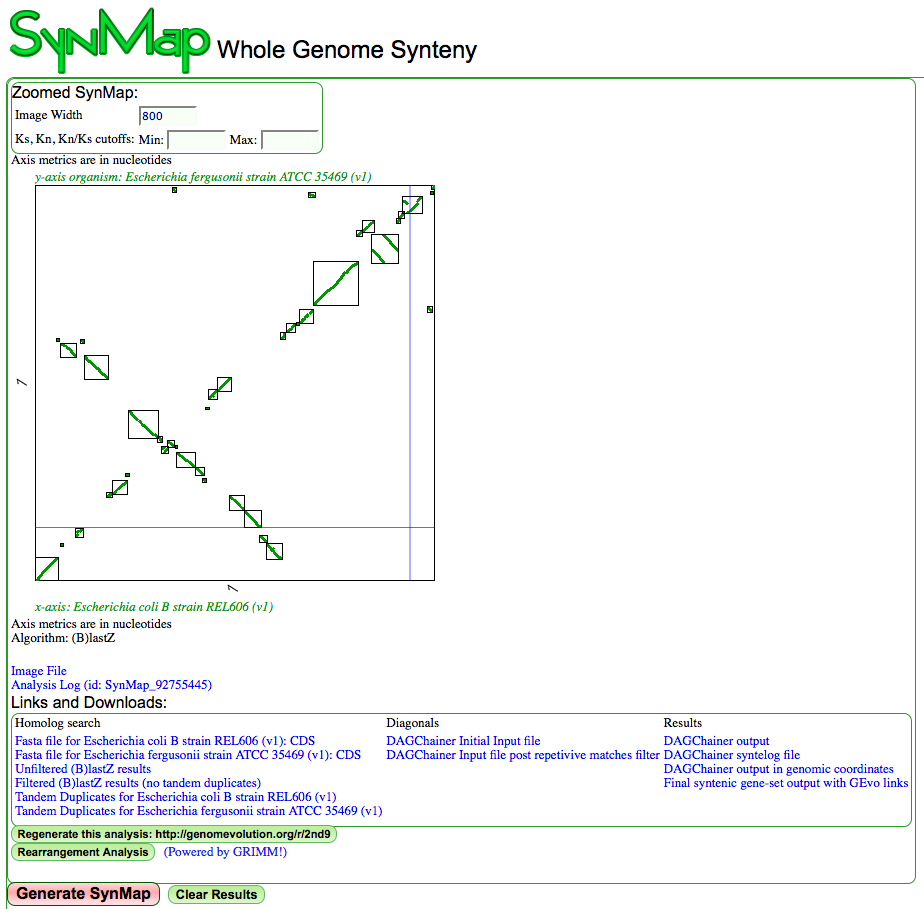
Syntentic dotplot between two species of Escherichia. X-axis: coli B strain REL606; Y-axis: fergusonii strain ATCC 35469. Syntenic dotplot has boxes drawn around syntenic regions and there is a link to GRIMM for rearrangement analysis. The link to GRIMM will only show if the quota align algorithm has been set and set to a 1:1 ratio. Results may be regenerated at http://genomevolution.org/r/2nd7
GRIMM auto-populated by SynMap
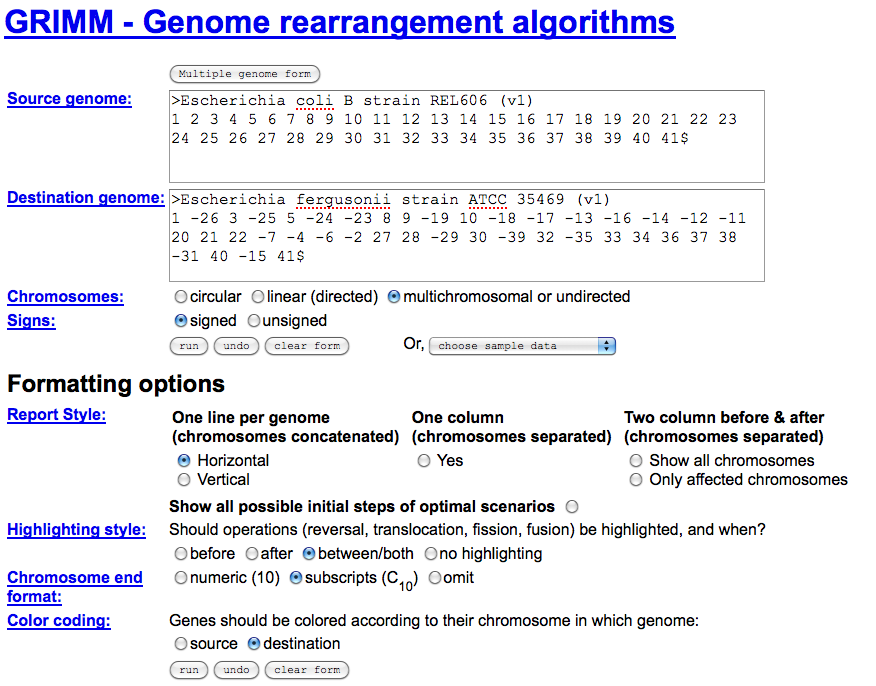
Screen-shot from GRIMM server with submission boxes auto-populated when linked to from SynMap
GRIMM genomic rearrangement analysis results

Screen-shot from GRIMM server after performing a rearrangement analysis on two Escherichia genomes.