Difference between revisions of "Plasmodia comparative genomics"
From CoGepedia
| Line 6: | Line 6: | ||
[[Image:Plasmodium-codon-substitution-matirx.png|thumb|right|600px|Log-odds score substitution matrix of codons between Plasmodium falciparum (x-axis) and Plasmodium knowlesi (y-axis). P. falciparum is a low-GC genome and P. kowlesi is a mid-GC genome.]] | [[Image:Plasmodium-codon-substitution-matirx.png|thumb|right|600px|Log-odds score substitution matrix of codons between Plasmodium falciparum (x-axis) and Plasmodium knowlesi (y-axis). P. falciparum is a low-GC genome and P. kowlesi is a mid-GC genome.]] | ||
| + | |||
| + | The genomes of two Plasmodium species, falciparum and knowlesi are structurally very similar to one another: | ||
| + | |||
| + | {| | ||
| + | |Organism || Chromosome count || Genome Length || CDS count || Genome GC content || CDS GC content || CDS Wobble position content || non-coding GC content | ||
| + | |Plasmodium falicparum || 14 || 22,860,235 bp || 5267 || 19.88% || 23.72% || 17.30% || 14.58% | ||
| + | |Plasmodium knowlesi || 14 || 23,462,187 bp || 5102 || 38.94% || 40.23% || 45.56% || 35.12% | ||
| + | } | ||
Revision as of 12:42, 29 March 2010

Syntenic dotplot of Plasmodium falciparum (x-axis) and Plasmodium knowlesi (y-axis). Results can be regenerated at: http://synteny.cnr.berkeley.edu/CoGe/SynMap.pl?dsgid1=7949;dsgid2=2465;c=4;D=20;g=10;A=5;Dm=0;gm=0;w=0;b=1;ft1=1;ft2=1;do1=1;do2=1;do=40;dt=geneorder
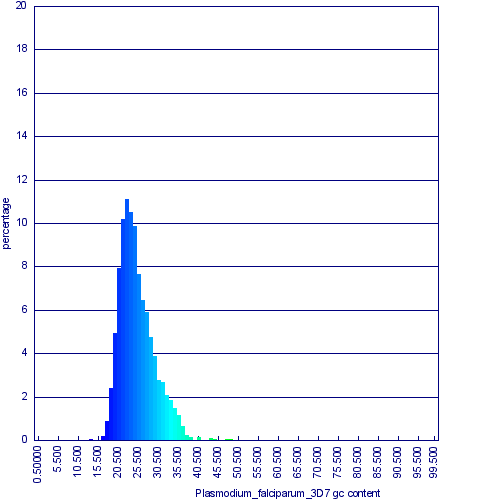
Histogram of CDS GC content for Plasmodium falciparum. Generated by OrganismView

Histogram of CDS GC content for Plasmodium knowlesi. Generated by OrganismView
The genomes of two Plasmodium species, falciparum and knowlesi are structurally very similar to one another:
| Organism | Chromosome count | Genome Length | CDS count | Genome GC content | CDS GC content | CDS Wobble position content | non-coding GC content | Plasmodium falicparum | 14 | 22,860,235 bp | 5267 | 19.88% | 23.72% | 17.30% | 14.58% | Plasmodium knowlesi | 14 | 23,462,187 bp | 5102 | 38.94% | 40.23% | 45.56% | 35.12%
} |
