Identifying SNPs
From CoGepedia
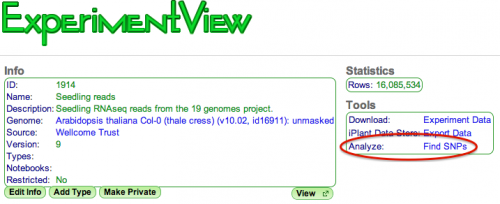
CoGe has a built-in SNP finding pipeline! To find SNPs in an existing alignment-type experiment, select the "Find SNPs" analysis tool in the ExperimentView page.
An alignment-type experiment in CoGe is created by loading a BAM file. See LoadExperiment for instruction on how to do this.
The CoGe SNP finding pipeline detects SNPs that meet the following criteria (these values are configurable):
- minimum read depth of 10 (of any base quality value)
- minimum high-quality allele count of 4 (high quality is PHRED >= 20)
- minimum allele frequency of 10%.
The SAMtools mpileup function is used to generate read depth counts for the BAM file.
We are working to incorporate other SNP-finding programs into CoGe. Let us know if you'd like your favorite tool added!