Inversion
From CoGepedia
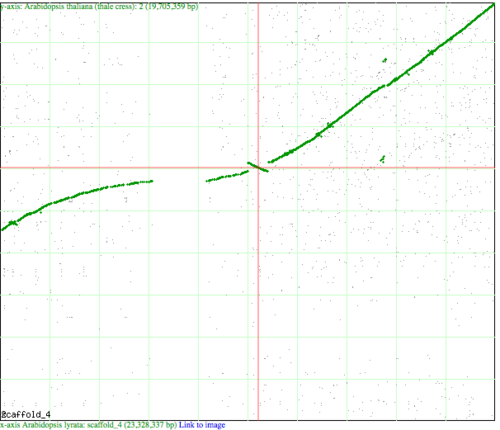
Syntenic dotplot visualized by SynMap between chromosome 4 of Arabidopsis lyrata (x-axis) and chromosome 2 of Arabidopsis thaliana (y-axis). Green dots represent syntenic genes as identified by DAGChainer. Note the inversion at the center of the cross hairs where the green "line" changes slope from positive to negative.

GEvo analysis of the inversion detected between the two Arabidopsis chromosomes. Results can be regenerated at http://toxic.berkeley.edu/CoGe/GEvo.pl?prog=blastz;spike_len=0;accn1=fgenesh2_kg.4__765__AT2G28060.1;fid1=30750039;dsid1=39129;dsgid1=3068;gstid1=1;chr1=scaffold_4;dr1up=830865;dr1down=1073591;gbstart1=1;gblength1=2332;mask1=non-cds;do1=1;accn2=AT2G28060;fid2=20219657;dsid2=35172;dsgid2=8;gstid2=1;chr2=2;dr2up=334450;dr2down=533528;gbstart2=1;gblength2=1591;mask2=non-cds;do2=2;num_seqs=2;hsp_overlap_limit=0;hsp_size_limit=0

Syntenic dotplot generated by SynMap between two strains of Yersinia pseudotuberculosis showing one inversion. Strain IP 31758 (x-axis); strain YPIII (y-axis). Results can be regenerated at: http://synteny.cnr.berkeley.edu/CoGe/SynMap.pl?dsgid1=1911;dsgid2=1906;D=20;g=10;A=5;w=0;b=1;ft1=1;ft2=1;dt=geneorder

GEvo analysis of inversion breakpoints identified in two strains of Yersinia pseudotuberculosis. At the breakpoints are ribosomal gene cassettes (shown as gray and yellow arrows). Results can be regenerated at: http://tinyurl.com/yj38xw6
Definition
A genomic inversion is when a region of a genome or chromosome gets flipped in place. The orientation of this genomic segment changes such that its 5' and 3' ends change places. These usually happen at genomic regions with nearly identical sequence, implying a mechanism similar to non-homologous recombination.
Bacteria
In bacterial genomes, inversion often occur at transposon or ribosomal gene sequences, and happen symmetrically around the origin of replication. The latter causes a characteristic pattern in syntenic dotplots called an x alignment.