this page is out of date. Please refer to EPIC-CoGe instead.
CoGe's genome visualization library, GeLo, is very flexible in the types of graphics it can produce. The best way to figure out what something means in an image from CoGe is to scroll through this list of images, find something that looks similar, and then read the description. Many of these visualization layers are available under the layers menu in GenomeView. If you are searching for a visualization layer that is not present in the layers menu, then that layer does not exist for that genome.
Overview
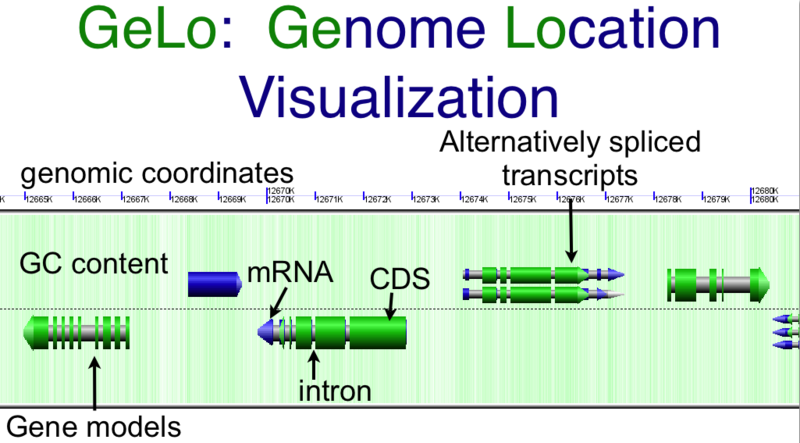
Overview of
GenomeView's visualization using the
GeLo genomic visualization library. Background of chromosome is colored green for GC rich regions and white for AT rich regions. Gene models are drawn as composite colored arrows such that gray is the extent of the gene, blue is mRNA, and green is protein coding
CDS sequence. Overlapping genomic features are detected and expanded to show alternatively spiced transcripts
Chromosome Background
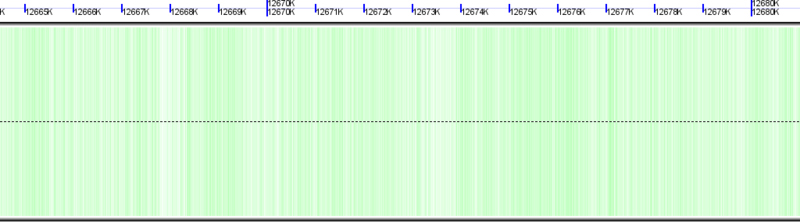
Background of chromosome colored green to show GC rich regions and white to show AT rich regions
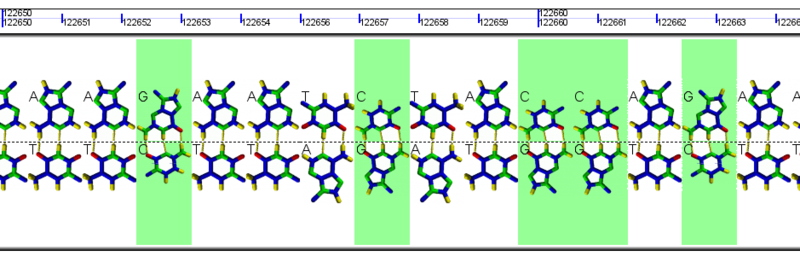
Example GeLo graphics using images of nucleotides in the background. These are also colored green for GC and white for AT
Genes
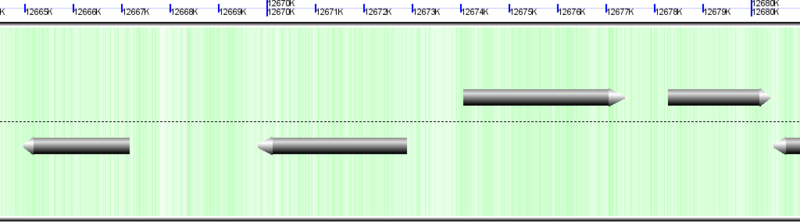
Gene models drawn as gray arrows
mRNAs
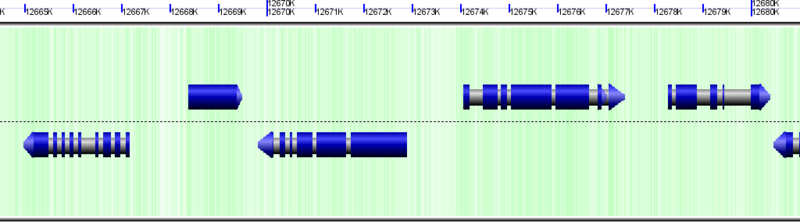
mRNA drawn as blue arrows (on top of gray genes)
CDSs
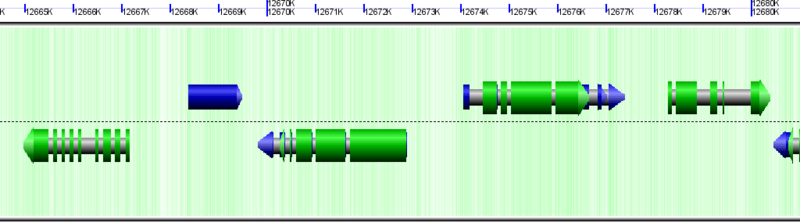
Protein coding regions (CDS) drawn as green arrows (on top of gray genes and blue mRNAs)
Unsequenced regions
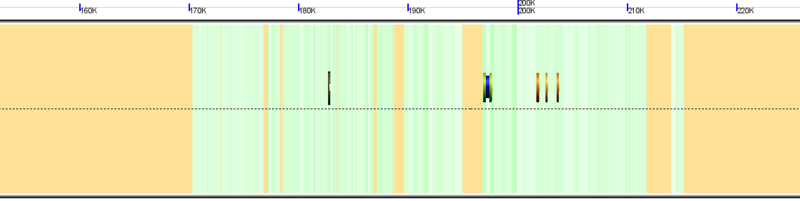
Genomic region with unsequenced regions colored orange from the genome of Carica papaya
Masked regions
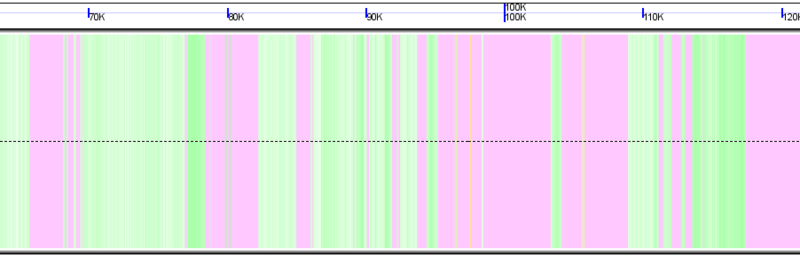
Genomic region with masked sequence colored purple
rRNA genes
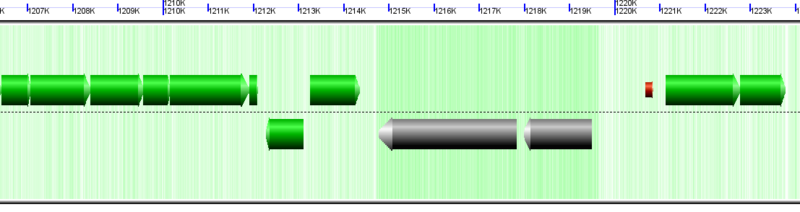
Microbial (archaea) genomic region with 23S and 16S rRNA genes drawn as wide gray arrows
Pseudogenes
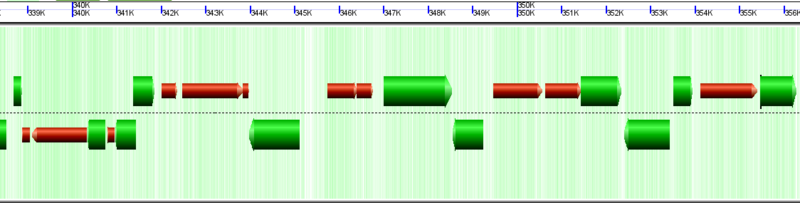
Microbial (archaea) genomic region with many pseudogenes drawn as orange-red arrows
Wobble GC content filter
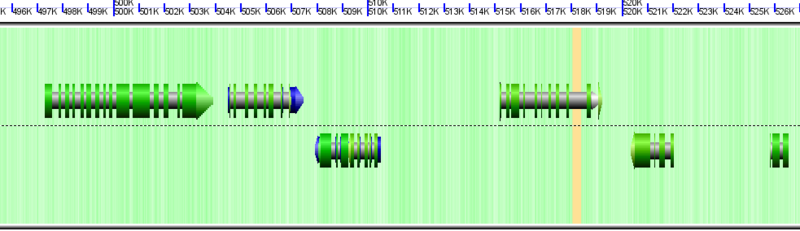
Wobble GC content colored for Chlamydomonas reinhardtii genome. Green means high GC content in codon wobble positions
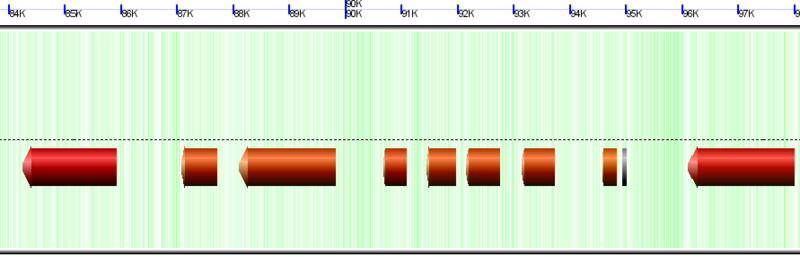
Wobble GC content colored for Chlamydomonas reinhardtii chloroplast genome. Red means low GC content in codon wobble positions
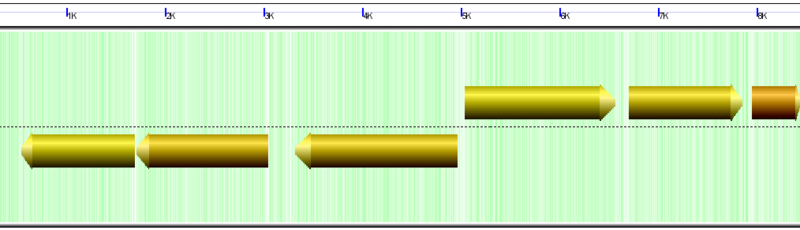
Wobble GC content colored for Chlamydomonas reinhardtii mitochondria genome. Yellow means ~50% GC content in codon wobble positions
GEvo Links
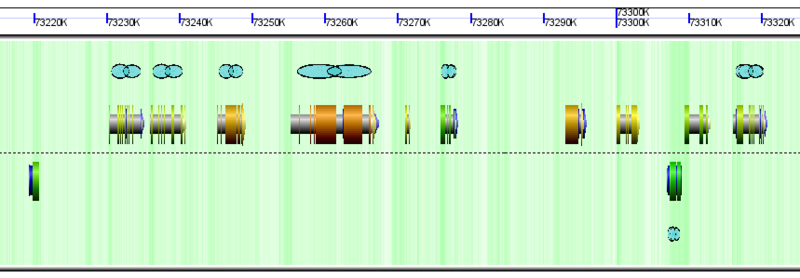
GenomeView showing glyphs for links to GEvo. These glyphs are blue "links" and are displayed when a genomic feature has a link to GEvo for syntenic analysis of syntelogs.
Tandem Duplicates
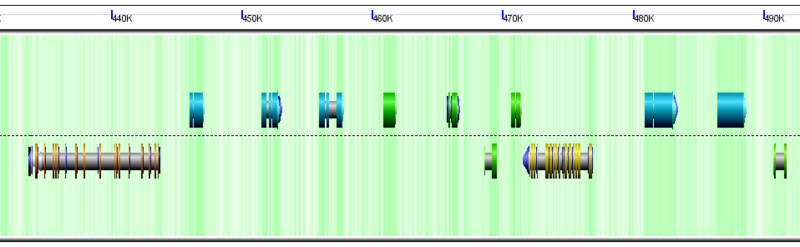
GenomeView showing tandem duplicate genes as blue arrows over
CDSs.
Repeat Regions
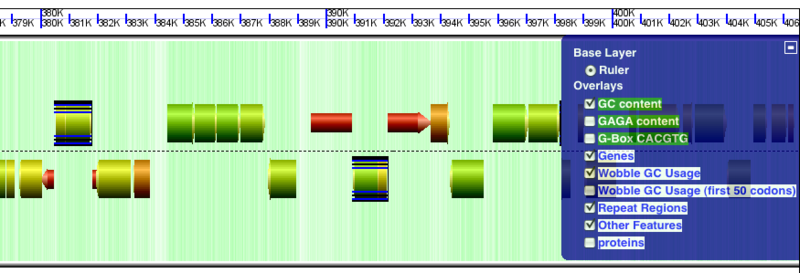
GenomeView showing repeat regions as black and blue boxes around a sequence and gene. There are two IS elements highlighted by these boxes that have each jumped into a gene and disrupted it.
Other Features
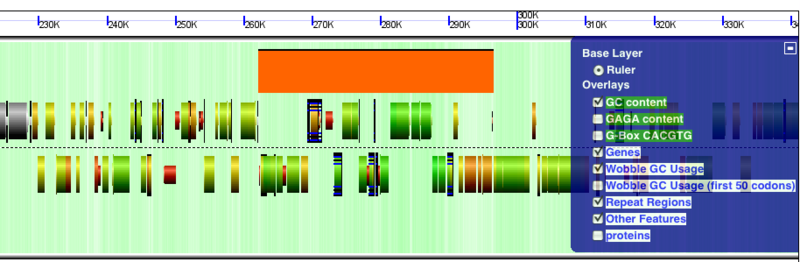
GenomeView showing "other features" as orange boxes. This other features is a prophage that has integrated in the genome of Escherichia coli K12 strain K-12 substrain MG1655.
















