GeLo
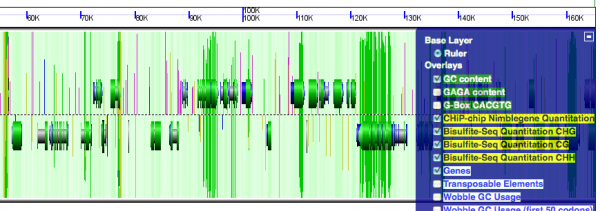
GeLo is CoGe's Genomic Visualization Library and stands for Genome Location Visualization. It is pronounced like the food-like item: Jello.
GeLo is used in various parts of CoGe for viewing genomic regions. One of the primary tools that uses it is GenomeView, which is CoGe's dynamic genome browser. Other tools in CoGe that use it are GEvo and CoGeBlast.
GeLo is similar to other genomic visualization libraries in the sense that it allows you to visualize a genomic region and paint genomic features on it using "tracks". However, GeLo differs substantially in how much freedom it provides to allow you to create many types of genomic visualization schemes. In essence, GeLo lets you create a virtual chromosome, paint it's background any way your wish, add genomic features to it based on nucleotide coodinates, and then will render your image for you based on the size you wish it to be. All the details of scaling the genomic features and determining their placement is handled for your behind the scenes. If the genomic feature is on the top strand, it is drawn in the top half of the chromosome automatically.
In terms of the control you have over the graphics you can:
- color the background of the chromosome independent of the stuff drawn on top of the background
- specify tracks for the placement of a genomic feature (track 1 is drawn closest to the middle, track 2 is drawn next closest, etc.)
- specify layers for a genomic features such that one feature is drawn on top of the another feature
- add labels to features
Also, you have the ability to create any type of image for a genomic feature glyph. This does require writing a perl module for it and using the GD library] for generating the glyph. However, you can use the full power of GD to generate your images including the ability to use other images, such as PNGs, in your glyph. This allows you to create some very interesting graphics such as using images of nucleotides.
See this example to see GeLo in action in CoGe's genome browser, GenomeView.
For more examples of different types of genomic visualization using GeLo see GenomeView examples.
You can download GeLo including an example implementation using CoGe's dynamic Genome Browser.

