Difference between revisions of "SeqView"
From CoGepedia
| Line 9: | Line 9: | ||
*Generating the reverse complement of the sequence | *Generating the reverse complement of the sequence | ||
*Translating the sequence | *Translating the sequence | ||
| − | *Extracting all [[genomic features]] encompassed by the sequence | + | *Extracting all [[genomic features]] encompassed by the sequence and sending them to [[FeatList]] |
*Calculating GC content of the sequence | *Calculating GC content of the sequence | ||
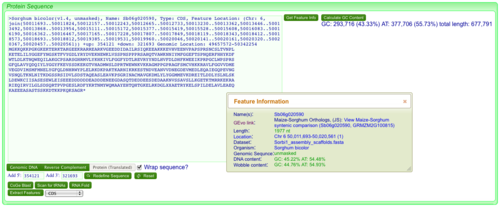
*Information about the feature used to anchoring the retrieved sequence (if applicable) | *Information about the feature used to anchoring the retrieved sequence (if applicable) | ||
Latest revision as of 21:51, 20 February 2010
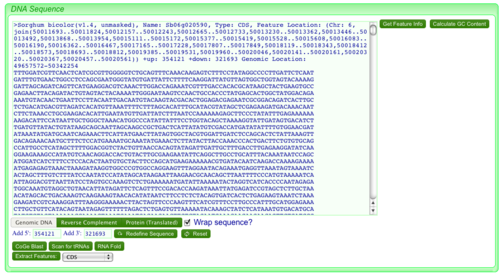
SeqView displays a single sequence in fasta format.
Features:
- Adding additional of 3' and 5' sequence
- Generating the reverse complement of the sequence
- Translating the sequence
- Extracting all genomic features encompassed by the sequence and sending them to FeatList
- Calculating GC content of the sequence
- Information about the feature used to anchoring the retrieved sequence (if applicable)
Links to:
- ViennaFold for determining RNA secondary structure
- CoGeBlast for searching genomes with the sequence
- tRNAView for searching the sequence for tRNAs